Researchers create a streamlined technique for generating proteins using E. coli
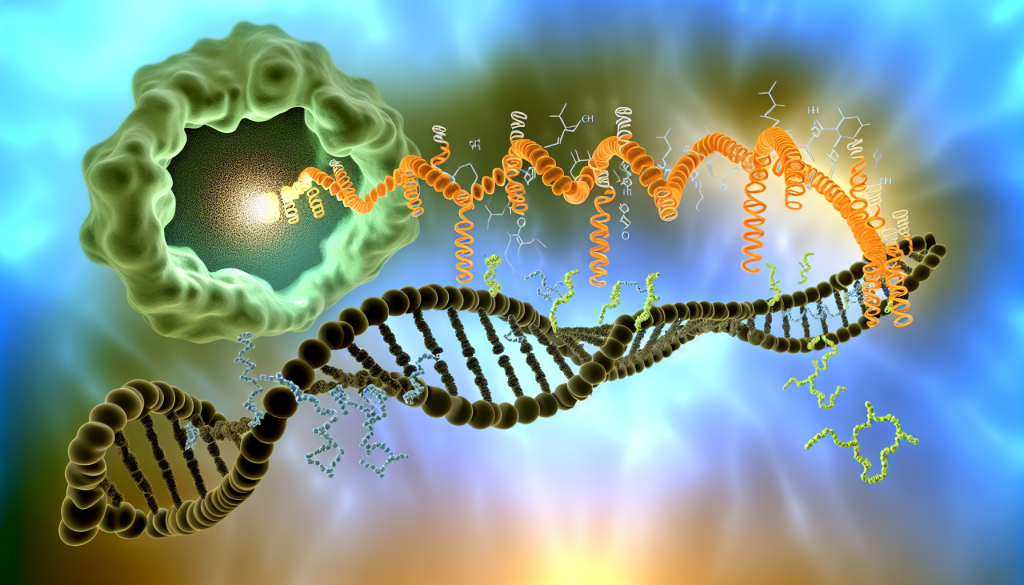
Image:
Proteins are created through two main steps involving DNA. First is transcription, where DNA is copied into mRNA. Then comes translation, during which ribosomes read the mRNA and join amino acids in the right order to form a protein. The illustration depicts a streamlined version of the translation process, enabling faster and more efficient protein production.
Credit: Teruyo Ojima-Kato
Microbial proteins are gaining interest for their usefulness in producing various products such as medicines, industrial enzymes, and antibodies used in diagnostics. They are also potential tools in converting renewable resources into biofuels and biodegradable plastics, offering alternatives to traditional fossil fuel-based products. As such, optimizing the way these proteins are created can support more eco-friendly manufacturing approaches.
Among microorganisms, Escherichia coli (E. coli) is widely used due to its cost-effectiveness and speed in protein production. However, the amount of protein produced can vary depending on the specific genetic code of the protein being synthesized. A team of scientists in Japan, reporting in the journal RSC Chemical Biology, has introduced a new method that enhances protein production in E. coli.
A common challenge is ribosome stalling, where ribosomes—the molecular machines that build proteins from mRNA—suddenly stop during the translation process. This halts protein production entirely. "In earlier work, we discovered that adding a very short peptide sequence made of four amino acids—serine, lysine, isoleucine, and lysine—at the beginning of a protein could reduce this stalling and improve how efficiently proteins are made in E. coli," explained Associate Professor Teruyo Ojima-Kato from Nagoya University's Graduate School of Bioagricultural Sciences.
Building on this concept, Professor Kato and Professor Hideo Nakano from Nagoya University, working with scientists from the National Institute of Advanced Industrial Science and Technology and Waseda University, began searching for other such peptides that could aid translation. They created a comprehensive tetrapeptide library, which includes every possible combination of four amino acids—totaling 160,000 different sequences.
Using this rich library, they identified several short peptides that effectively prevent ribosome stalling. The next challenge was evaluating the potential of all 160,000 tetrapeptides. To manage such a large-scale task, the researchers developed an artificial intelligence (AI) model trained on data from around 250 different experiments. After three rounds of predictions and testing, the AI accurately forecasted which peptides would enhance the translation process.
"Our study introduces a new strategy for boosting protein production using compact peptide sequences," said Kato. "This method could enhance the production of key enzymes needed in the bio-based chemical and fuel industry. Ultimately, our approach could form the basis of technologies for sustainable manufacturing that reduce dependence on fossil fuels."
Journal
RSC Chemical Biology
Article Publication Date
22-Oct-2025

